|
Time_point_grp < - split(cl_vertex$names[cl_vertex$last_mutation != G_mod < - igraph::graph_from_data_frame(d = cl_edges, directed = TRUE, Labels_show = labels_show, label_offset = label_offset)Ĭl_vertex < - cl_df$cl_vertex[order(cl_df$cl_vertex$TP, cl_df$cl_vertex$coord.y), Xc = Xc, Yc = Yc, cl_df = cl_df, adjMatrix_base = adjMatrix_base,ĪdjMatrix_overall = adjMatrix_overall, mut_TP = mut_TP, Idx_next_v < - which(colSums(adjMatrix_overall != 0) = 0)Ĭl_df < - recursiveLongitudinalLayout(idx_Vc = idx_next_v, Mut_TP < - cumsum(mut_TP) + 1:length(mut_TP)Ĭl_df < - list(cl_vertex = cl_vertex, cl_edges = cl_edges) 
Max_deep_tp < - max(max_deep_tp, deep_tmp) Ind_lPath = 1, mut_list = NA, row_v = idx)ĭeep_tmp < - recursiveDescend(descend)$max_deep Idx_anc_c < - which(colSums(adjMatrix_base_tp) =ĭescend = list(adjM = adjMatrix_base_tp, l_path = 0, Storage.mode(adjMatrix_overall) < - "integer"Ĭol_in < - which(colnames(adjMatrix_base) %in% cl_edges$name[cl_edges$to %in%ĪdjMatrix_base_tp < - adjMatrix_base G < - igraph::graph_from_data_frame(cl_edges, directed = TRUE,ĪdjMatrix_overall < - igraph::get.adjacency(g, sparse = F,ĪdjMatrix_overall < - 0ĪdjMatrix_overall < - 2ĪdjMatrix_overall < - 1 Igraph::delete_edge_attr(graph = g, name = "extincion") Igraph::vertex_attr(graph = g, name = "color") < - cl_vertex$color Igraph::vertex_attr(graph = g, name = "branch_level") < - cl_vertex$branch_level Igraph::vertex_attr(graph = g, name = "branch") < - cl_vertex$branch N_levels < - max(cl_vertex$branch_level < - "#C0C0C0"Ĭl_vertex$color < - "#DDDDDD" Parental_clones < - c(parental_clones, c_son) Igraph::write_graph(data, file = con, format = "graphml")Ĭat(paste0("Error writing: ", path, "\n", e))Ĭat(paste0(toupper(type), " file written: "))Ĭat(paste0("File unable to be written.\n"))ġ9 File: visualization.R, author: BIMIB-DISCo, license: Apache License 2.0 
Paste0(supported_types, collapse = ", "), "\n"))Ĭat(paste0("Error writing rds: ", path, "\n", e))Ĭon < - file(path, open = "wb", encoding = "native.enc") Supported_types < - c("graphml", "csv", "rds")Ĭat(paste0("File output not supported. Write_output_file < - function(data, type = "rds", name = "File", It will save the file in your working directory, or you can specify a different file path with the file argument.19 File: Utils.R, author: vosonlab, license: GNU General Public License v3.0 You simply specify which graph to save (called fig in our example) and specify the file location, name, and type with the file argument. The simplest method is to use the export function from the plotly package. These methods can be integrated into a loop or other programmatic process to easily create multiple graphs at once. You can also save plotly graphics with R code. You can share these files with colleagues so that they can open it in a web browser and interact with the graphic. This will save the graphic as an html file. 
That is, they don’t have any of the interactive features that make plotly so great! To retain those features, you can select the “Save as a web page…” option the Export menu. 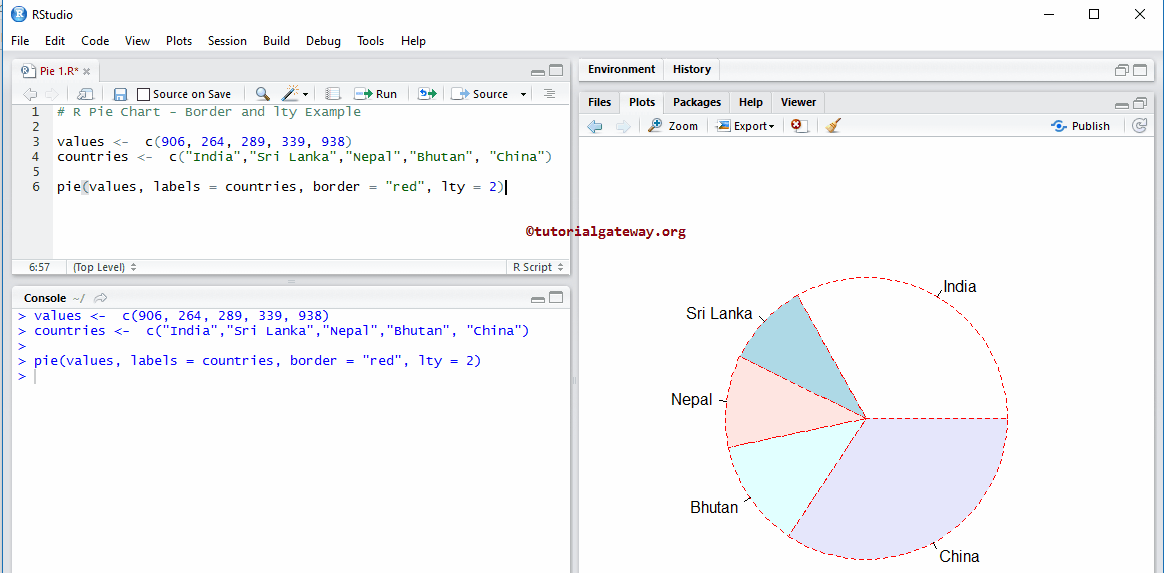
If you select “Save an image…” you can specify the image type (i.e., png, jpeg) and size before saving.īoth of these methods save a static image of the graph. If you are using RStudio, the “Export” menu will give you more control over the output file. When a plotly graph is displayed (either in the viewer window of RStudio or on a webpage), you can hover your cursor over the top of the graph to find the button here: Perhaps the easiest way to write your graphic as a PNG file is to click the “download plot as PNG” button in the toolbar at the top of the graph.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
